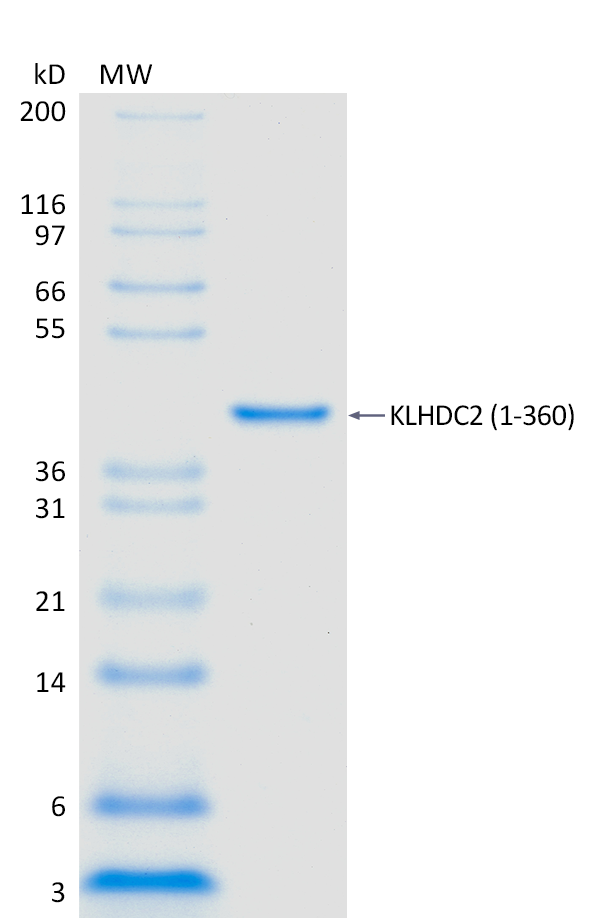
 2 μg KLHDC2 run on 4-12% SDS-PAGE gel under reducing conditions, then visualized with Colloidal Coomassie Blue Stain.
2 μg KLHDC2 run on 4-12% SDS-PAGE gel under reducing conditions, then visualized with Colloidal Coomassie Blue Stain.For Research Use Only (RUO)
Scott D.C., et al. (2023) Mol Cell 83(5): 770-786. PMID: 36805027
Hickey C.H., et al. (2024) Nat Struct Mol Biol 31(2):311-322. PMID: 38177675