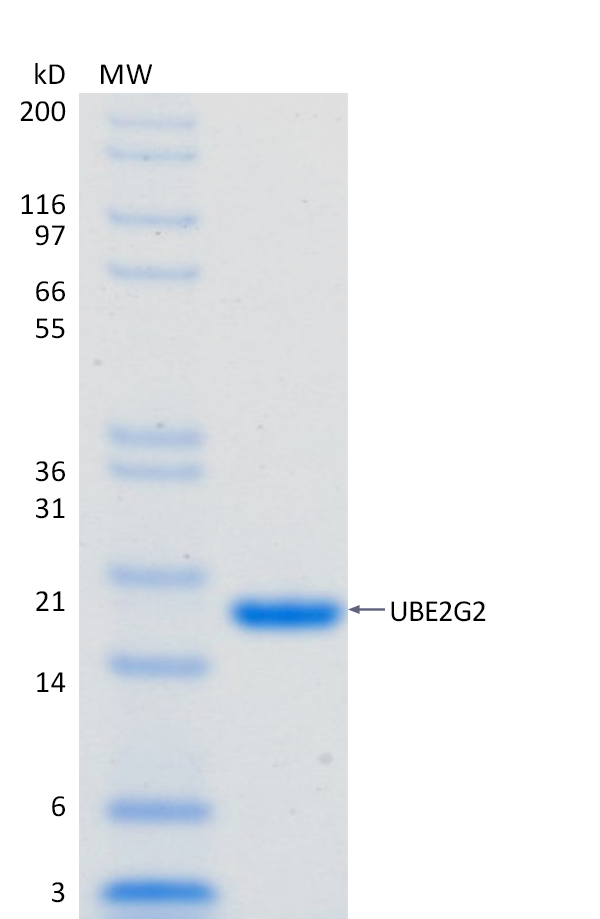
 2 μg UBE2G2 run on 4-12% SDS-PAGE gel under reducing conditions, then visualized with Colloidal Coomassie Blue Stain.
2 μg UBE2G2 run on 4-12% SDS-PAGE gel under reducing conditions, then visualized with Colloidal Coomassie Blue Stain.For Research Use Only (RUO)
Tsai P-L., et al. (2022) Cell Reports 41(8):111675. PMID 36417855
Roussel B.D., et al. (2013) Hum Mol Genet 22(22):4616-4626. PMID 23814041